Electrochemical gradient

An electrochemical gradient is a gradient of electrochemical potential, usually for an ion that can move across a membrane. The gradient consists of two parts:
- The chemical gradient, or difference in solute concentration across a membrane.
- The electrical gradient, or difference in charge across a membrane.
If there are unequal concentrations of an ion across a permeable membrane, the ion will move across the membrane from the area of higher concentration to the area of lower concentration through simple diffusion. Ions also carry an electric charge that forms an electric potential across a membrane. If there is an unequal distribution of charges across the membrane, then the difference in electric potential generates a force that drives ion diffusion until the charges are balanced on both sides of the membrane.
Electrochemical gradients are essential to the operation of batteries and other electrochemical cells, photosynthesis and cellular respiration, and certain other biological processes.
Overview
[edit]Electrochemical energy is one of the many interchangeable forms of potential energy through which energy may be conserved. It appears in electroanalytical chemistry and has industrial applications such as batteries and fuel cells. In biology, electrochemical gradients allow cells to control the direction ions move across membranes. In mitochondria and chloroplasts, proton gradients generate a chemiosmotic potential used to synthesize ATP,[1] and the sodium-potassium gradient helps neural synapses quickly transmit information.[citation needed]
An electrochemical gradient has two components: a differential concentration of electric charge across a membrane and a differential concentration of chemical species across that same membrane. In the former effect, the concentrated charge attracts charges of the opposite sign; in the latter, the concentrated species tends to diffuse across the membrane to an equalize concentrations. The combination of these two phenomena determines the thermodynamically-preferred direction for an ion's movement across the membrane.[2]: 403 [3]
The combined effect can be quantified as a gradient in the thermodynamic electrochemical potential:[citation needed]
with
- μi the chemical potential of the ion species i
- zi the charge per ion of the species i
- F, Faraday constant (the electrochemical potential is implicitly measured on a per-mole basis)
- φ, the local electric potential.
Sometimes, the term "electrochemical potential" is abused to describe the electric potential generated by an ionic concentration gradient; that is, φ.
An electrochemical gradient is analogous to the water pressure across a hydroelectric dam. Routes unblocked by the membrane (e.g. membrane transport protein or electrodes) correspond to turbines that convert the water's potential energy to other forms of physical or chemical energy, and the ions that pass through the membrane correspond to water traveling into the lower river.[tone] Conversely, energy can be used to pump water up into the lake above the dam, and chemical energy can be used to create electrochemical gradients.[4][5]
Chemistry
[edit]The term typically applies in electrochemistry, when electrical energy in the form of an applied voltage is used to modulate the thermodynamic favorability of a chemical reaction. In a battery, an electrochemical potential arising from the movement of ions balances the reaction energy of the electrodes. The maximum voltage that a battery reaction can produce is sometimes called the standard electrochemical potential of that reaction.
Biological context
[edit]The generation of a transmembrane electrical potential through ion movement across a cell membrane drives biological processes like nerve conduction, muscle contraction, hormone secretion, and sensation. By convention, physiological voltages are measured relative to the extracellular region; a typical animal cell has an internal electrical potential of (−70)–(−50) mV.[2]: 464
An electrochemical gradient is essential to mitochondrial oxidative phosphorylation. The final step of cellular respiration is the electron transport chain, composed of four complexes embedded in the inner mitochondrial membrane. Complexes I, III, and IV pump protons from the matrix to the intermembrane space (IMS); for every electron pair entering the chain, ten protons translocate into the IMS. The result is an electric potential of more than 200 mV. The energy resulting from the flux of protons back into the matrix is used by ATP synthase to combine inorganic phosphate and ADP.[6][2]: 743–745
Similar to the electron transport chain, the light-dependent reactions of photosynthesis pump protons into the thylakoid lumen of chloroplasts to drive the synthesis of ATP. The proton gradient can be generated through either noncyclic or cyclic photophosphorylation. Of the proteins that participate in noncyclic photophosphorylation, photosystem II (PSII), plastiquinone, and cytochrome b6f complex directly contribute to generating the proton gradient. For each four photons absorbed by PSII, eight protons are pumped into the lumen.[2]: 769–770
Several other transporters and ion channels play a role in generating a proton electrochemical gradient. One is TPK3, a potassium channel that is activated by Ca2+ and conducts K+ from the thylakoid lumen to the stroma, which helps establish the electric field. On the other hand, the electro-neutral K+ efflux antiporter (KEA3) transports K+ into the thylakoid lumen and H+ into the stroma, which helps establish the pH gradient.[7]
Ion gradients
[edit]
Since the ions are charged, they cannot pass through cellular membranes via simple diffusion. Two different mechanisms can transport the ions across the membrane: active or passive transport.[citation needed]
An example of active transport of ions is the Na+-K+-ATPase (NKA). NKA is powered by the hydrolysis of ATP into ADP and an inorganic phosphate; for every molecule of ATP hydrolized, three Na+ are transported outside and two K+ are transported inside the cell. This makes the inside of the cell more negative than the outside and more specifically generates a membrane potential Vmembrane of about −60 mV.[5]
An example of passive transport is ion fluxes through Na+, K+, Ca2+, and Cl− channels. Unlike active transport, passive transport is powered by the arithmetic sum of osmosis (a concentration gradient) and an electric field (the transmembrane potential). Formally, the molar Gibbs free energy change associated with successful transport is[citation needed] where R represents the gas constant, T represents absolute temperature, z is the charge per ion, and F represents the Faraday constant.[2]: 464–465
In the example of Na+, both terms tend to support transport: the negative electric potential inside the cell attracts the positive ion and since Na+ is concentrated outside the cell, osmosis supports diffusion through the Na+ channel into the cell. In the case of K+, the effect of osmosis is reversed: although external ions are attracted by the negative intracellular potential, entropy seeks to diffuse the ions already concentrated inside the cell. The converse phenomenon (osmosis supports transport, electric potential opposes it) can be achieved for Na+ in cells with abnormal transmembrane potentials: at +70 mV, the Na+ influx halts; at higher potentials, it becomes an efflux.[citation needed]
| Ion | Mammal | Squid axon | S. cerevisiae | E. coli | Sea water | ||
|---|---|---|---|---|---|---|---|
| Cell | Blood | Cell | Blood | ||||
| K+ | 100 - 140 | 4-5 | 400 | 10 - 20 | 300 | 30 - 300 | 10 |
| Na+ | 5-15 | 145 | 50 | 440 | 30 | 10 | 500 |
| Mg2+ | 10 [a] 0.5 - 0.8 [b] |
1 - 1.5 | 50 | 30 - 100 [a] 0.01 - 1 [b] |
50 | ||
| Ca2+ | 10−4 | 2.2 - 2.6 [c] 1.3 - 1.5 [d] |
10−4 - 3×10−4 | 10 | 2 | 3 [a] 10−4 [b] |
10 |
| Cl− | 4 | 110 | 40 - 150 | 560 | 10 - 200 [e] | 500 | |
| X− (negatively charged proteins) | 138 | 9 | 300 - 400 | 5-10 | |||
| HCO3− | 12 | 29 | |||||
| pH | 7.1 - 7.3[12] | 7.35 to 7.45 [12] (normal arterial blood pH) 6.9 - 7.8 [12] (overall range) |
7.2 - 7.8[13] | 8.1 - 8.2[14] | |||
Proton gradients
[edit]Proton gradients in particular are important in many types of cells as a form of energy storage. The gradient is usually used to drive ATP synthase, flagellar rotation, or metabolite transport.[15] This section will focus on three processes that help establish proton gradients in their respective cells: bacteriorhodopsin and noncyclic photophosphorylation and oxidative phosphorylation.[citation needed]
Bacteriorhodopsin
[edit]
The way bacteriorhodopsin generates a proton gradient in Archaea is through a proton pump. The proton pump relies on proton carriers to drive protons from the side of the membrane with a low H+ concentration to the side of the membrane with a high H+ concentration. In bacteriorhodopsin, the proton pump is activated by absorption of photons of 568nm wavelength, which leads to isomerization of the Schiff base (SB) in retinal forming the K state. This moves SB away from Asp85 and Asp212, causing H+ transfer from the SB to Asp85 forming the M1 state. The protein then shifts to the M2 state by separating Glu204 from Glu194 which releases a proton from Glu204 into the external medium. The SB is reprotonated by Asp96 which forms the N state. It is important that the second proton comes from Asp96 since its deprotonated state is unstable and rapidly reprotonated with a proton from the cytosol. The protonation of Asp85 and Asp96 causes re-isomerization of the SB, forming the O state. Finally, bacteriorhodopsin returns to its resting state when Asp85 releases its proton to Glu204.[15][16]
Photophosphorylation
[edit]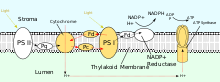
PSII also relies on light to drive the formation of proton gradients in chloroplasts, however, PSII utilizes vectorial redox chemistry to achieve this goal. Rather than physically transporting protons through the protein, reactions requiring the binding of protons will occur on the extracellular side while reactions requiring the release of protons will occur on the intracellular side. Absorption of photons of 680nm wavelength is used to excite two electrons in P680 to a higher energy level. These higher energy electrons are transferred to protein-bound plastoquinone (PQA) and then to unbound plastoquinone (PQB). This reduces plastoquinone (PQ) to plastoquinol (PQH2) which is released from PSII after gaining two protons from the stroma. The electrons in P680 are replenished by oxidizing water through the oxygen-evolving complex (OEC). This results in release of O2 and H+ into the lumen, for a total reaction of[15] Failed to parse (syntax error): {\displaystyle 4h\nu+2\ce{H2O}+2\ce{PQ}+4\ce{H+}(\text{stroma})\longrightarrow\ce{O2}+2\ce{PQH2}+4\ce{H+}(\text{lumen})}
After being released from PSII, PQH2 travels to the cytochrome b6f complex, which then transfers two electrons from PQH2 to plastocyanin in two separate reactions. The process that occurs is similar to the Q-cycle in Complex III of the electron transport chain. In the first reaction, PQH2 binds to the complex on the lumen side and one electron is transferred to the iron-sulfur center which then transfers it to cytochrome f which then transfers it to plastocyanin. The second electron is transferred to heme bL which then transfers it to heme bH which then transfers it to PQ. In the second reaction, a second PQH2 gets oxidized, adding an electron to another plastocyanin and PQ. Both reactions together transfer four protons into the lumen.[2]: 782–783 [17]
Oxidative phosphorylation
[edit]
In the electron transport chain, complex I (CI) catalyzes the reduction of ubiquinone (UQ) to ubiquinol (UQH2) by the transfer of two electrons from reduced nicotinamide adenine dinucleotide (NADH) which translocates four protons from the mitochondrial matrix to the IMS:[18] Failed to parse (syntax error): {\displaystyle \ce{NADH} + \ce{H^+} + \ce{UQ} + 4\underbrace{\ce{H^+}}_{\mathrm{matrix}} \longrightarrow \ce{NAD^+} + \ce{UQH_2} + 4\underbrace{\ce{H^+}}_{\mathrm{IMS}}}
Complex III (CIII) catalyzes the Q-cycle. The first step involving the transfer of two electrons from the UQH2 reduced by CI to two molecules of oxidized cytochrome c at the Qo site. In the second step, two more electrons reduce UQ to UQH2 at the Qi site. The total reaction is:[18] Failed to parse (syntax error): {\displaystyle 2\underbrace{\text{cytochrome c}}_{\text{oxidized}}+\ce{UQH_2}+2\underbrace{\ce{H^+}}_{\text{matrix}}\longrightarrow2\underbrace{\text{cytochrome c}}_{\text{reduced}}+\ce{UQ}+4\underbrace{\ce{H^+}}_{\text{IMS}}}
Complex IV (CIV) catalyzes the transfer of two electrons from the cytochrome c reduced by CIII to one half of a full oxygen. Utilizing one full oxygen in oxidative phosphorylation requires the transfer of four electrons. The oxygen will then consume four protons from the matrix to form water while another four protons are pumped into the IMS, to give a total reaction[18] Failed to parse (syntax error): {\displaystyle 2\text{cytochrome c}(\text{reduced})+4\ce{H+}(\text{matrix})+\frac{1}{2}\ce{O2}\longrightarrow2\text{cytochrome c}(\text{oxidized})+2\ce{H+}(\text{IMS})+\ce{H2O}}
See also
[edit]References
[edit]- ^ Nath, Sunil; Villadsen, John (2015-03-01). "Oxidative phosphorylation revisited". Biotechnology and Bioengineering. 112 (3): 429–437. doi:10.1002/bit.25492. ISSN 1097-0290. PMID 25384602. S2CID 2598635.
- ^ a b c d e f Nelson, David; Cox, Michael (2013). Lehninger Principles of Biochemistry. New York: W.H. Freeman. ISBN 978-1-4292-3414-6.
- ^ Yang, Huanghe; Zhang, Guohui; Cui, Jianmin (2015-01-01). "BK channels: multiple sensors, one activation gate". Frontiers in Physiology. 6: 29. doi:10.3389/fphys.2015.00029. PMC 4319557. PMID 25705194.
- ^ Shattock, Michael J.; Ottolia, Michela; Bers, Donald M.; Blaustein, Mordecai P.; Boguslavskyi, Andrii; Bossuyt, Julie; Bridge, John H. B.; Chen-Izu, Ye; Clancy, Colleen E. (2015-03-15). "Na+/Ca2+ exchange and Na+/K+-ATPase in the heart". The Journal of Physiology. 593 (6): 1361–1382. doi:10.1113/jphysiol.2014.282319. ISSN 1469-7793. PMC 4376416. PMID 25772291.
- ^ a b Aperia, Anita; Akkuratov, Evgeny E.; Fontana, Jacopo Maria; Brismar, Hjalmar (2016-04-01). "Na+-K+-ATPase, a new class of plasma membrane receptors". American Journal of Physiology. Cell Physiology. 310 (7): C491–C495. doi:10.1152/ajpcell.00359.2015. ISSN 0363-6143. PMID 26791490.
- ^ Poburko, Damon; Demaurex, Nicolas (2012-04-24). "Regulation of the mitochondrial proton gradient by cytosolic Ca2+ signals" (PDF). Pflügers Archiv: European Journal of Physiology. 464 (1): 19–26. doi:10.1007/s00424-012-1106-y. ISSN 0031-6768. PMID 22526460. S2CID 18133149.
- ^ Höhner, Ricarda; Aboukila, Ali; Kunz, Hans-Henning; Venema, Kees (2016-01-01). "Proton Gradients and Proton-Dependent Transport Processes in the Chloroplast". Frontiers in Plant Science. 7: 218. doi:10.3389/fpls.2016.00218. PMC 4770017. PMID 26973667.
- ^ Philips, Ron Milo & Ron. "» What are the concentrations of different ions in cells?". Retrieved 2019-06-07.
- ^ Lodish, Harvey; Berk, Arnold; Zipursky, S. Lawrence; Matsudaira, Paul; Baltimore, David; Darnell, James (2000). "Table 15-1, Typical Ion Concentrations in Invertebrates and Vertebrates". www.ncbi.nlm.nih.gov. Retrieved 2019-06-07.
- ^ "The following table gives an idea of the intra and extra cellular ion concentrations in a squid axon and a mammalian cell". www.chm.bris.ac.uk. Retrieved 2019-06-07.
- ^ Diem K, Lenter C. Scientific Tables. Vol. 565 (Seventh ed.). Basel: Ciba-Geigy Limited. pp. 653–654. ISBN 978-3-9801244-0-9.
- ^ a b c Spitzer, Kenneth W.; Vaughan-Jones, Richard D. (2003), Karmazyn, Morris; Avkiran, Metin; Fliegel, Larry (eds.), "Regulation of Intracellular pH in Mammalian Cells", The Sodium-Hydrogen Exchanger: From Molecule to its Role in Disease, Springer US, pp. 1–15, doi:10.1007/978-1-4615-0427-6_1, ISBN 9781461504276
- ^ Slonczewski, Joan L.; Wilks, Jessica C. (2007-08-01). "pH of the Cytoplasm and Periplasm of Escherichia coli: Rapid Measurement by Green Fluorescent Protein Fluorimetry". Journal of Bacteriology. 189 (15): 5601–5607. doi:10.1128/JB.00615-07. ISSN 0021-9193. PMC 1951819. PMID 17545292.
- ^ Brewer, Peter G. (September 1, 2008). "Rising Acidity in the Ocean: The Other CO2 Problem". doi:10.1038/scientificamericanearth0908-22.
{{cite journal}}: Cite journal requires|journal=(help) - ^ a b c Gunner, M. R.; Amin, Muhamed; Zhu, Xuyu; Lu, Jianxun (2013-08-01). "Molecular mechanisms for generating transmembrane proton gradients". Biochimica et Biophysica Acta (BBA) - Bioenergetics. Metals in Bioenergetics and Biomimetics Systems. 1827 (8–9): 892–913. doi:10.1016/j.bbabio.2013.03.001. PMC 3714358. PMID 23507617.
- ^ Wickstrand, Cecilia; Dods, Robert; Royant, Antoine; Neutze, Richard (2015-03-01). "Bacteriorhodopsin: Would the real structural intermediates please stand up?". Biochimica et Biophysica Acta (BBA) - General Subjects. Structural biochemistry and biophysics of membrane proteins. 1850 (3): 536–553. doi:10.1016/j.bbagen.2014.05.021. PMID 24918316.
- ^ Schöttler, Mark Aurel; Tóth, Szilvia Z.; Boulouis, Alix; Kahlau, Sabine (2015-05-01). "Photosynthetic complex stoichiometry dynamics in higher plants: biogenesis, function, and turnover of ATP synthase and the cytochrome b 6 f complex". Journal of Experimental Botany. 66 (9): 2373–2400. doi:10.1093/jxb/eru495. ISSN 0022-0957. PMID 25540437.
- ^ a b c Sun, Fei; Zhou, Qiangjun; Pang, Xiaoyun; Xu, Yingzhi; Rao, Zihe (2013-08-01). "Revealing various coupling of electron transfer and proton pumping in mitochondrial respiratory chain". Current Opinion in Structural Biology. 23 (4): 526–538. doi:10.1016/j.sbi.2013.06.013. PMID 23867107.
- Campbell & Reece (2005). Biology. Pearson Benjamin Cummings. ISBN 978-0-8053-7146-8.
- Stephen T. Abedon, "Important words and concepts from Chapter 8, Campbell & Reece, 2002 (1/14/2005)", for Biology 113 at the Ohio State University


